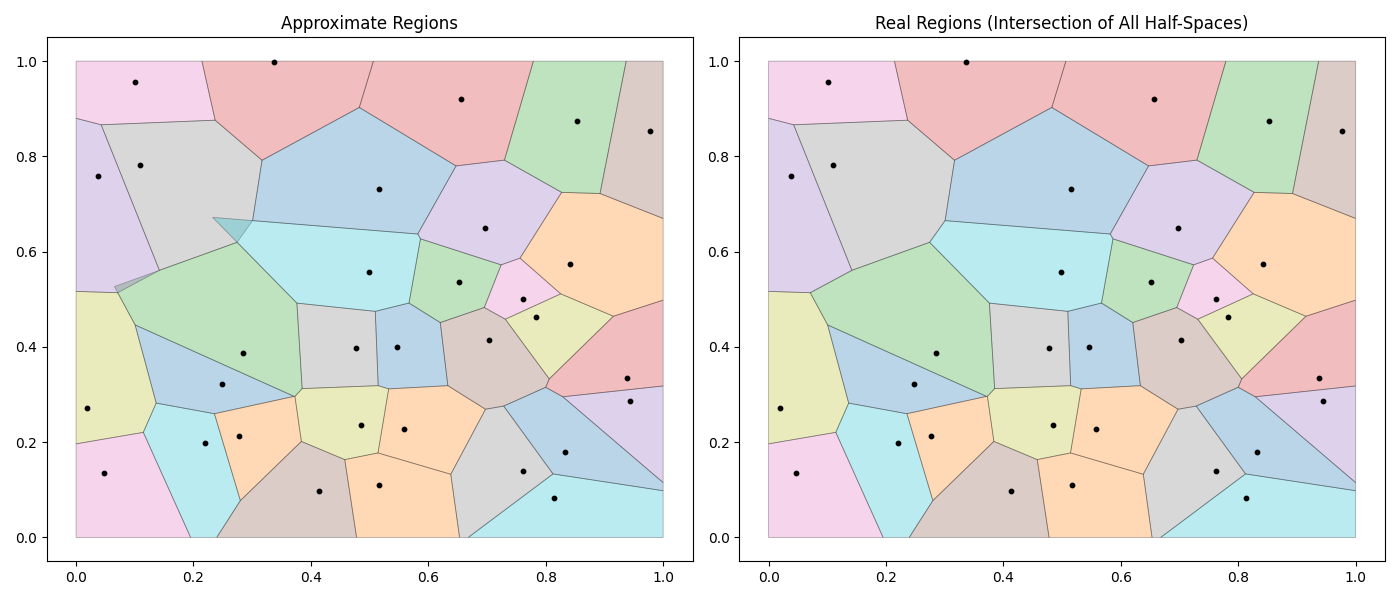

Earlier this week, I came up with an algorithm for generating approximate voronoi regions. It’s not optimal in most scenarios, but it has some nice properties:
Consider a set of sites . We’ll store all sites in a standard quadtree; this will give us inserts and deletes and range queries. To serve queries quickly, Bowyer-Watson maintains multiple data structures to store triangulation edges and vertices. By only storing sites, our algorithm saves a good chunk of memory at the cost of needing to compute the voronoi regions on-the-fly. Determine for yourself if this is worthwhile…
For in a metric space , the voronoi region for any site can be computed with a series of intersections. For simplicity, we will proceed using and Euclidean distance. With these assumptions, for each pair of sites and , we can partition into two half-spaces and split along the perpendicular bisector of the midpoint of . We call the side closer to than .
However, it is not necessary to compute for each of sites. The only relevant sites are the direct “neighbors” of . It has been shown that in , regardless of , the number of “neighbors” for a site is (see: notes on planar graphs). To return accurate voronoi regions, it’s sufficient to compute on the set of neighbors, . If we select an extra non-neighbor into , doesn’t affect on the resulting region and excluding a neighbor from will result in a region that is a superset of the true region. The problem can now be reframed as “how can we select an that includes all ‘neighbors’ with high probability?”.
— We don’t actually know how the distribution of “neighbors” evolves as increases. Some simulation-based work has shown values for , beyond that, information is very sparse. See simulation results - fig. 10.
I claim that if we define as the sites nearest to , we can produce an accurate voronoi region . Inserting a new site to the tree is and computing requires a range query. Overall, this operation is , and we can always recompute the region with another call to reduce overlap.
— If you read the code, you will notice that in the main text I’ve omitted the complexity of the iterative clipping step. In theory, this is as each clip could add an edge. I’m hand-waving a bit here, but because we expect a constant number of “neighbors”, we also expect a constant number of vertices at each step. I believe it is safe to treat this operation as . Ask yourself, do I care about the asymptotic complexity? or do we care about the constants? This isn’t a pretty algorithm, so I assume you’re here for the constants…
The cost of a delete depends on how important it is to preserve . I have mixed thoughts on this part:
If we do not care about full coverage, this operation is just a quadtree delete. If we must fill the resulting void, we’ll need to recompute for some set of nearby cells.
Assuming sites are drawn at random, I think recomputing the regions of the nearest neighbors succeeds . In this scheme, a delete is equivalent to inserts and has cost .
Recomputing the nearest neighbors may not be sufficient if the distribution of sites changes. As an alternative, I propose that when a region is first computed, we store the length of the region along its dominant axis. At delete time, we can construct a conservative estimate of the region’s bounding box and recompute the regions for all overlapping sites. This operation is equivalent to a constant number of inserts.
— We must be a bit careful with the asymptotic complexities. If the density of the cell is increasing, then a delete operation can involve many sites.
I benchmarked my implementation (with a slight assist from *️⃣) on datasets of 16M and 32M sites and found we can handle up to operations/s. This is quite slow, but perhaps it’s good enough to keep sites in memory and get just OK performance on region generation.
Benchmark_Voronoi/parallel-size=16M_neighbors=3ln(n)-8 180944 6843 ns/op
Benchmark_Voronoi/baseline-size=16M_neighbors=3ln(n)-8 52060 23019 ns/op
Benchmark_Voronoi/parallel-size=32M_neighbors=3ln(n)-8 73656 26218 ns/op
Benchmark_Voronoi/baseline-size=32M_neighbors=3ln(n)-8 24957 50666 ns/opI ran the following program with 32M points and found it used just under 800MB. This is actually quite nice compared to the required to store sites and triangulations. Personally, if I were running some sort of large-scale logistics operation that required tracking 32M entities, I’d opt for the Bowyer algorithm and a cloud bill that’s /month higher, but this is just my preference…
Showing nodes accounting for 796.53MB, 100% of 796.53MB total
flat flat% sum% cum cum%
528.02MB 66.29% 66.29% 528.02MB 66.29% github.com/paulmach/orb/quadtree.(*Quadtree).add
268.51MB 33.71% 100% 796.53MB 100% main.(*VoronoiGraph).Add
0 0% 100% 528.02MB 66.29% github.com/paulmach/orb/quadtree.(*Quadtree).Add
0 0% 100% 796.53MB 100% main.mainfunc main() {
go http.ListenAndServe("localhost:6060", nil)
done := make(chan bool)
bl, tr := orb.Point{0, 0}, orb.Point{1, 1}
vg := NewVoronoiGraph(orb.Bound{Min: bl, Max: tr})
rng := rand.New(rand.NewSource(rand.Int63n(1000)))
for i := 0; i < 32 * 1024 * 1024; i++ {
f64x, f64y := rng.Float64(), rng.Float64()
_ = vg.Add(f64x, f64y, 0)
}
for _, nn := range vg.tree.InBound(nil, orb.Bound{bl, tr}) {
pt := nn.Point()
id := idFromCoord(pt[0], pt[1])
_ = vg.ApproxVoronoiCell(id, 2.0, true, nil)
}
<-done
}Some final notes:
I didn’t discuss it at all, but depends on clipping according to a variation of the Sutherland–Hodgman algorithm. This was not my focus this week, and a better implementation of this algorithm can probably cut execution time by a good deal.
As always, I’m stubbornly refusing to use Github Gist, so the code is available here.